Alignment
Visualizes the sequence alignment of your antibodies using standard numbering schemes.
This tool is useful for evaluating differences between sequences and for generating IP claims around functional positions.
Accessing the Tool
Select at least one antibody in the Project View. Go to the Analysis menu and select Alignment. This will open the Alignment workspace in a new tab.
Variable Region
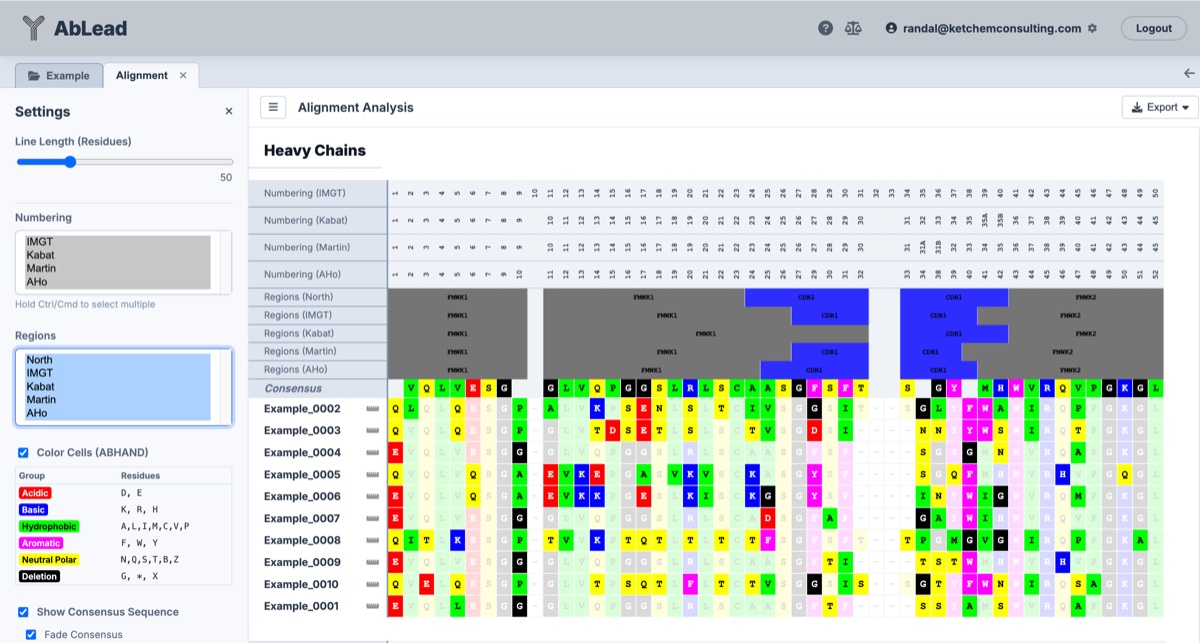
Constant Region
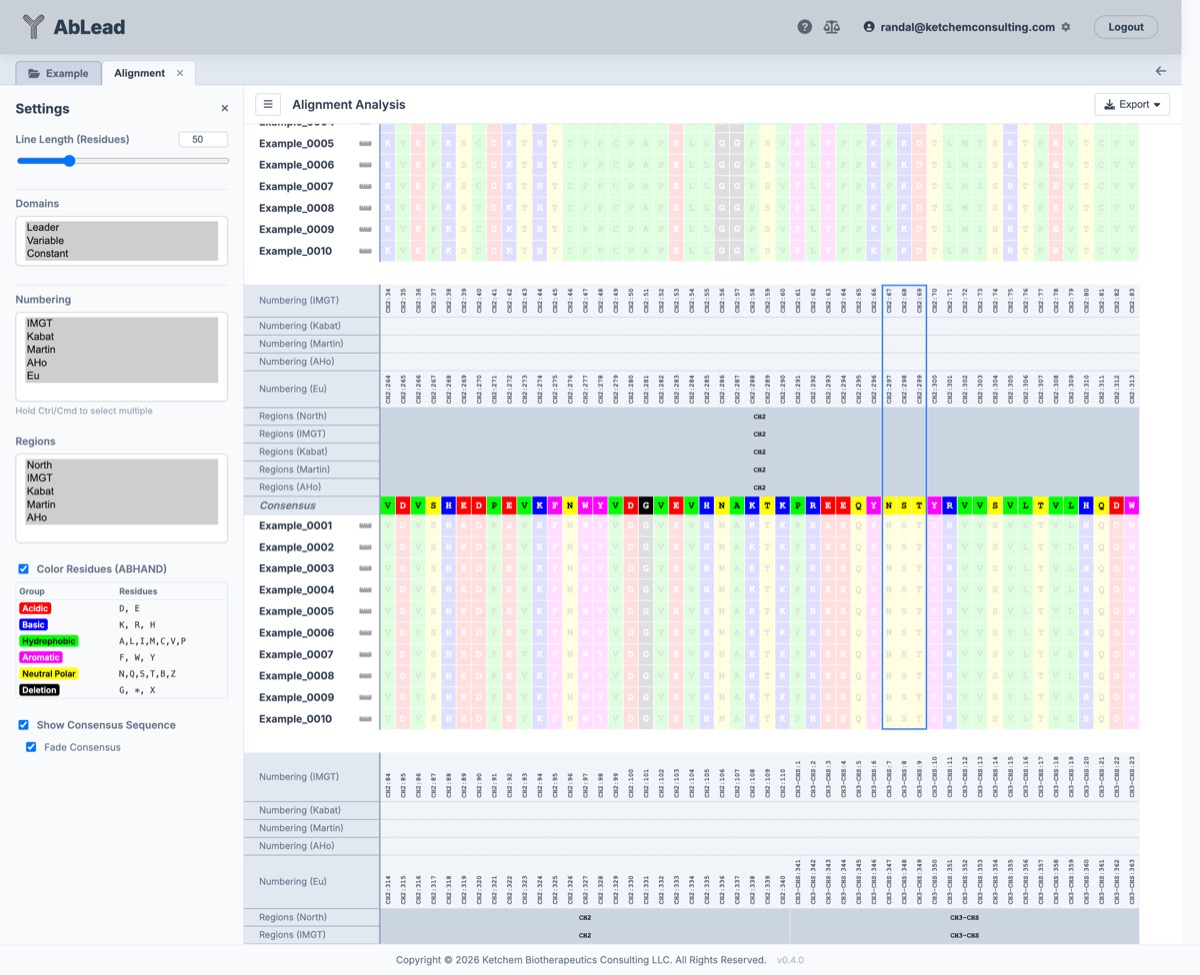
Using the Tool
- Grouping: Sequences are grouped by chain type (Kappa, Lambda, Heavy).
- Settings
- Domains: The Leader, Variable, and Constant regions may be displayed.
- Numbering and Regions: The numbering and region scheme may be changed by selecting different schemes from their respective selection boxes.
- Leader sequences are numbered negatively from the start of the variable domain.
- Constant domains are numbered based on an alignment of the germline sequences for that region.
- Color Residues: The residues may be colored or not based on ABHAND.
- Consensus: A consensus sequence may be displayed at the top of each group.
- Fade Consensus: Toggle this option to gray out residues that match the consensus, highlighting mutations and unique features.
- Linear Sequence: Click the icon next to the entry name to toggle the linear and mature linear number displays above a sequence.
- Export
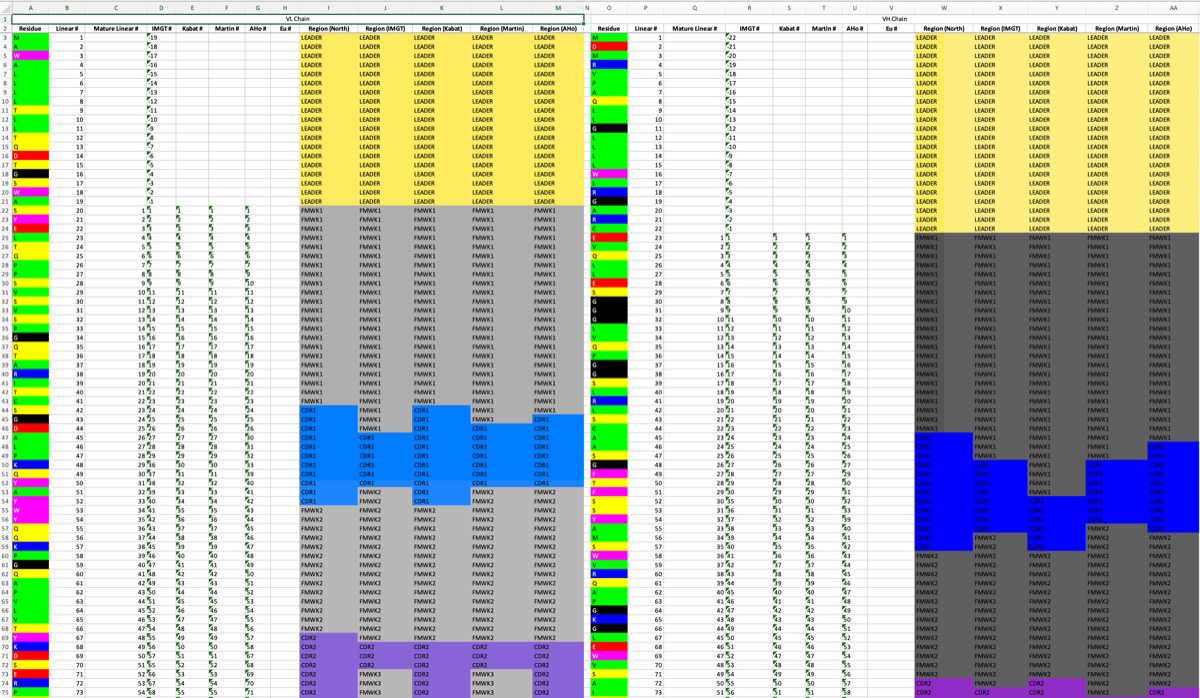
- Export Numbering Sheet: Exports the sequences with all numbering and region depictions.
- Export Alignment Excel: Exports the alignment in an Excel file.
- Engineering Mutation Designs: Click a residue to enter a mutation design used by the Engineering tab.
- Observations: Click a residue to enter an observation for that IMGT position. These are displayed in the Observations tab and on a clicked residue.
- Reorder Sequences: Click and drag the grip icon on the left side of a sequence row to reorder the sequences within their chain group. Note that this will also reorder the entry in the chain group for the other chain.
- Indicator Dots: Dots on a residue position indicate that there is an associated engineering design or observation at that position. Engineering mutations will appear as a small eggshell square while observations will appear as a small orange circle.
Excel Export
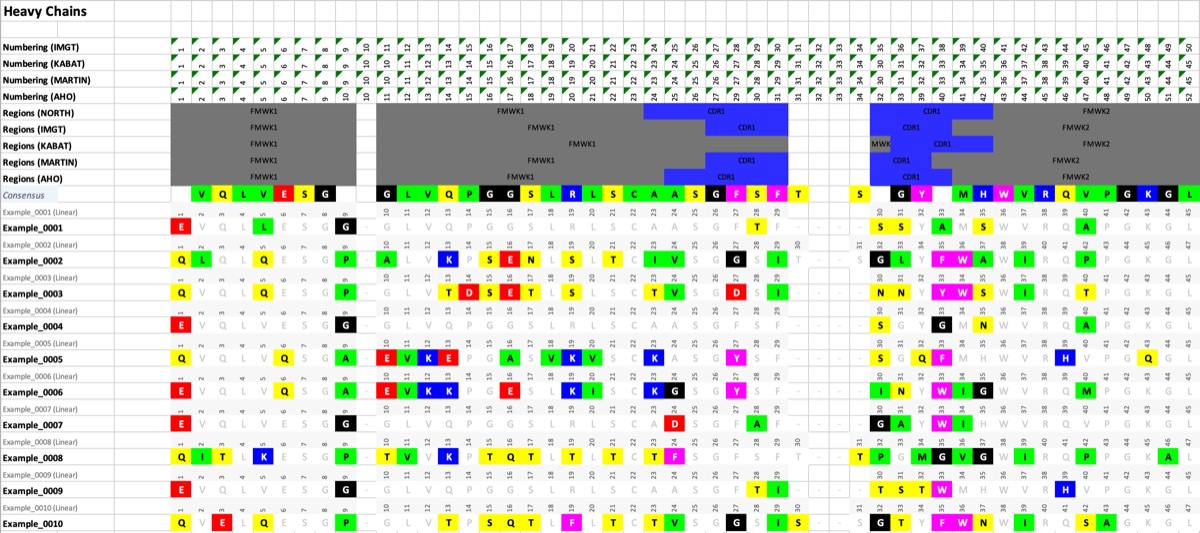

Numbering Sheet
Numbering Sheet Set

Numbering Sheet Entry